Selected work
Featured Research


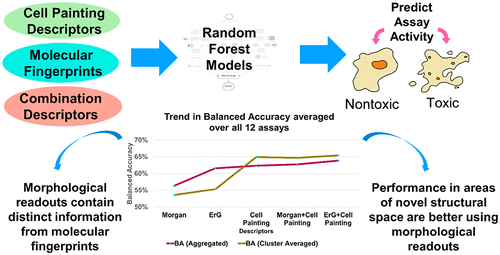
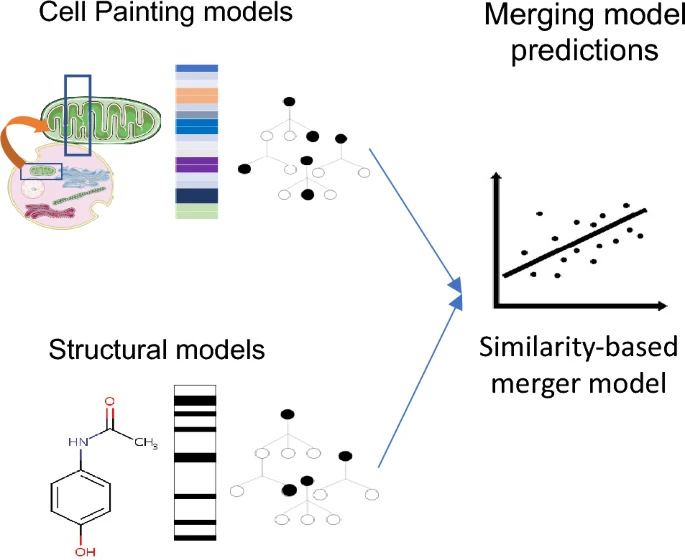
Cell Painting for cytotoxicity and mode-of-action analysis in primary human hepatocytes
Read paper
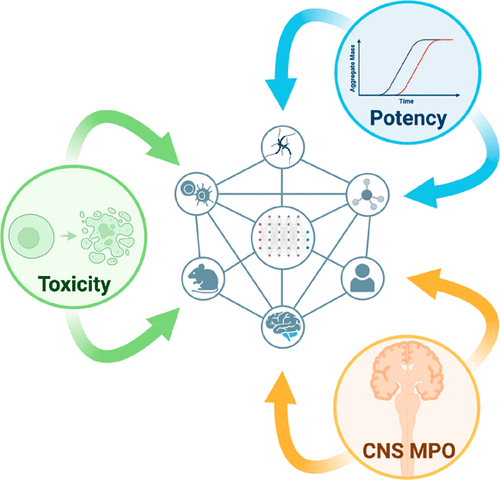
PKSmart: an open-source computational model to predict intravenous pharmacokinetics of small molecules
Read paper